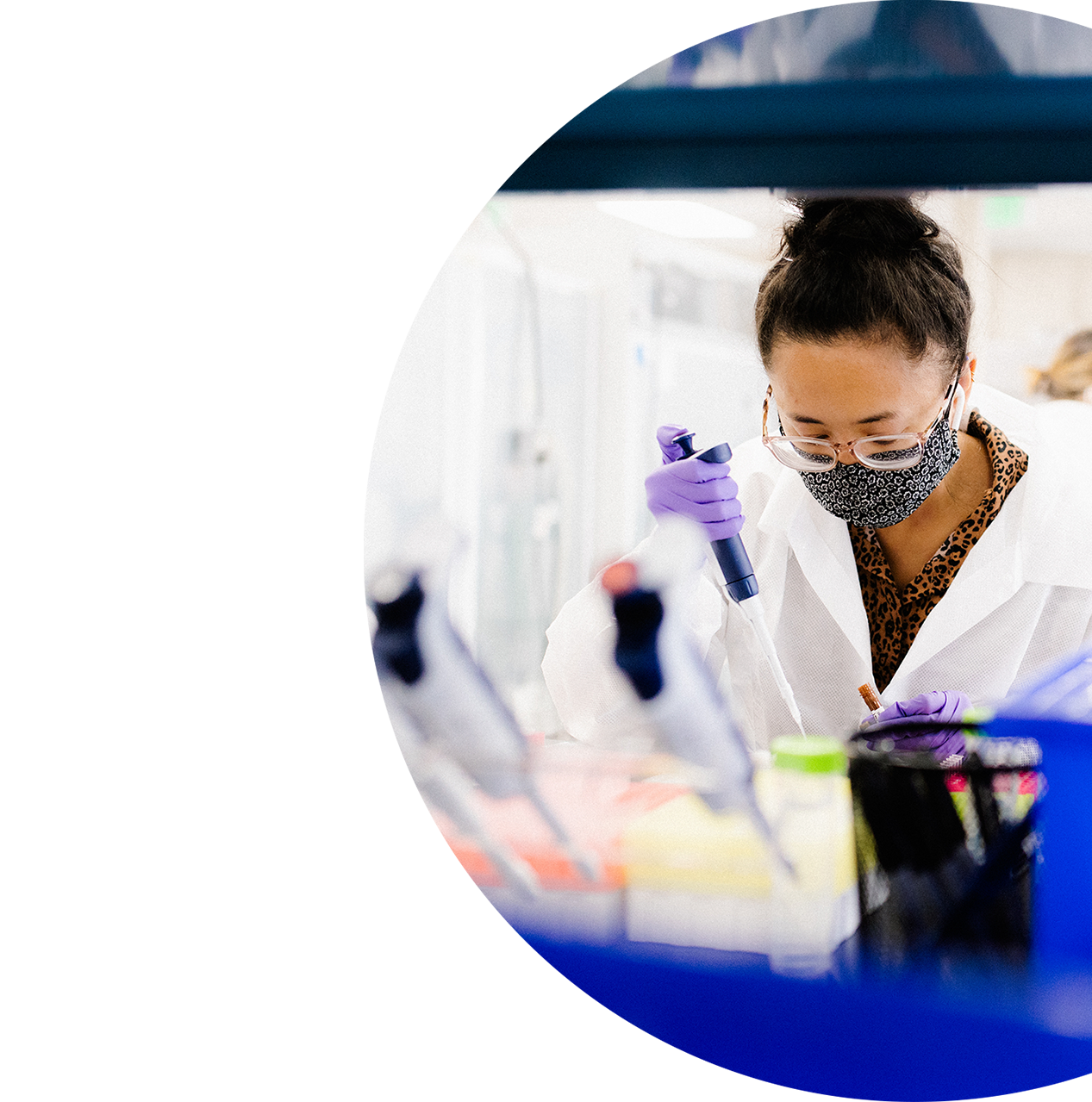
Blood Cancer Research
Deep molecular profiling of blood cancers
Blood cancer research tests detect various driver mutations in hematologic malignancies, such as myeloid and lymphoid diseases, by targeted next-generation sequencing (NGS).
This approach combines FusionPlex® and VariantPlex® panels to characterize gene fusions, point mutations, copy number variations (CNVs) and other variant types from a single sample.

Key Features
Unified
Workflow
Our panels are powered by Anchored Multiplex PCR (AMP™) chemistry and follow the same workflow for maximum efficiency

Comprehensive Tumor
Profiling
Get the most information from your sample - fusions, CNVs, SNVs, indels and expression levels.
Complex Variant
Detection
Coverage combined with powerful bioinformatics enable variant detection of traditionally difficult regions like CEBPA & FLT3
Confident Variant
Calling
Powerful bioinformatics combined with orthogonal variant verification.
RNA Input
Input
Reads
Genes
DNA Input
DNA
Reads
Genes
